Gut bacteriophage found inside Bacteroides fragilis links to colorectal cancer in metagenomic cohort
Intriguing signal but causality and confounding still do the heavy lifting
Images
 Data: Colorectal cancer is one of the deadliest forms for people under 50
foxnews.com
Data: Colorectal cancer is one of the deadliest forms for people under 50
foxnews.com
 Medical illustration of Colorectal Cancer
foxnews.com
Medical illustration of Colorectal Cancer
foxnews.com
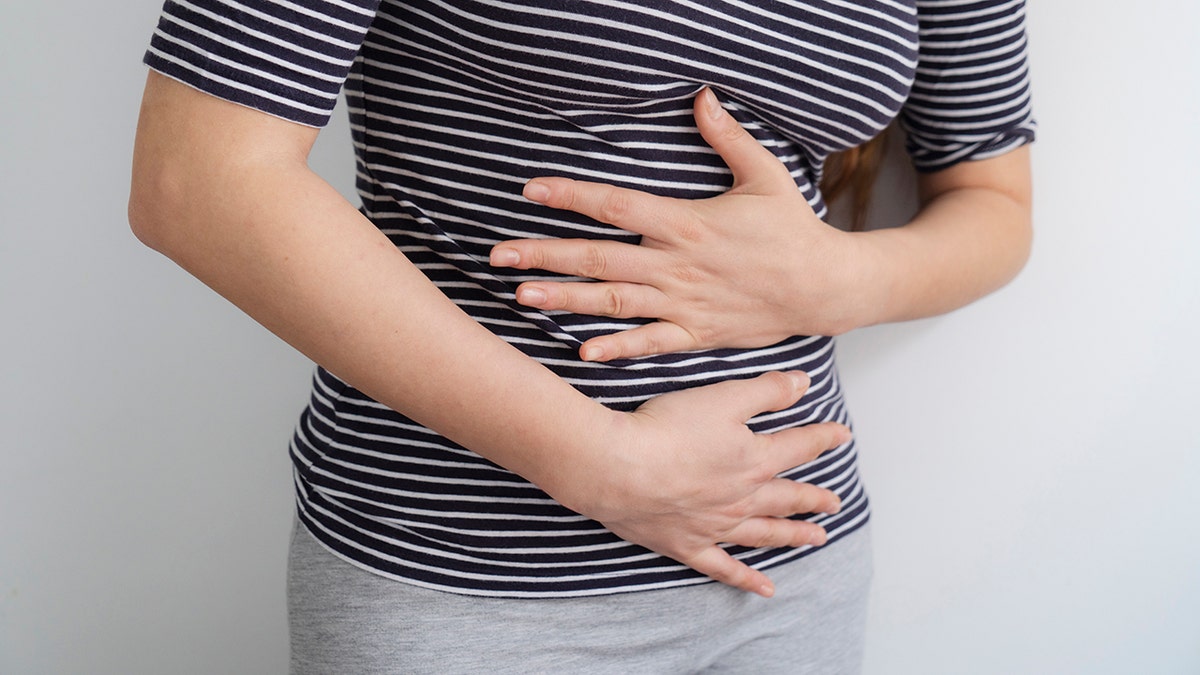 Woman's midsection seen as she clutches her abdomen in pain.
foxnews.com
Woman's midsection seen as she clutches her abdomen in pain.
foxnews.com
 Scientist working with test scopes and microscope in lab.
foxnews.com
Scientist working with test scopes and microscope in lab.
foxnews.com
 Male doctor using large intestine model with colonoscopy exam report on computer screen is explaining about colorectal cancer status to patient elderly man sitting at desk across from him.
foxnews.com
Male doctor using large intestine model with colonoscopy exam report on computer screen is explaining about colorectal cancer status to patient elderly man sitting at desk across from him.
foxnews.com
 Sarah and her mother, Jo (Family handout)
Family handout
Sarah and her mother, Jo (Family handout)
Family handout
 Sarah died in her father Mark’s arms last year (Family handout)
Family handout
Sarah died in her father Mark’s arms last year (Family handout)
Family handout
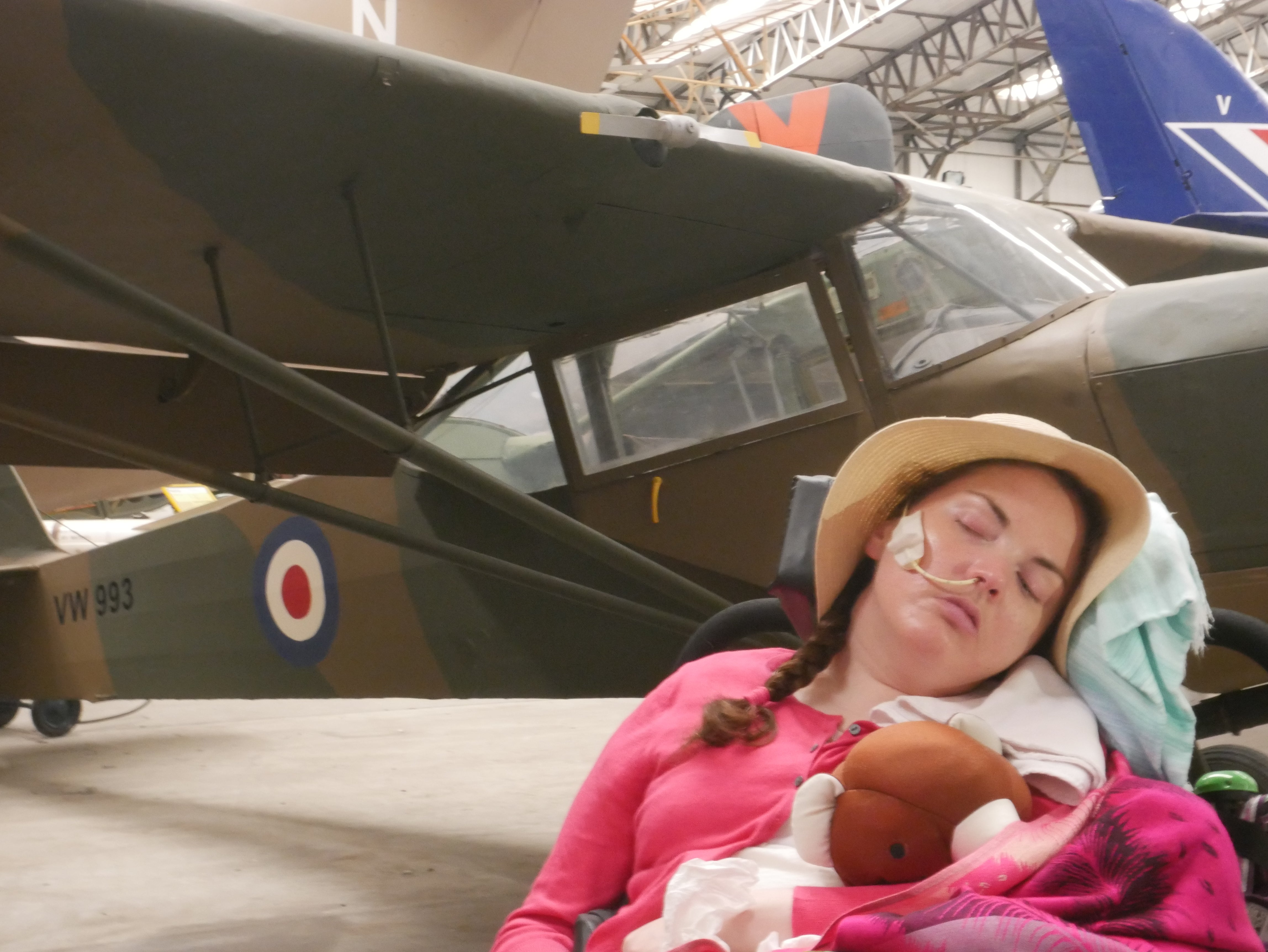 Sarah Walton (Family handout)
Family handout
Sarah Walton (Family handout)
Family handout
A newly described bacteriophage—essentially a virus that infects bacteria rather than human cells—has been associated with roughly double the odds of colorectal cancer in a multi-cohort stool metagenomics analysis, according to researchers at Odense University Hospital and the University of Southern Denmark.
Fox News reports that the team detected the phage embedded within strains of Bacteroides fragilis, a common gut commensal that has long been observed more frequently in colorectal cancer patients. That older association has been hard to interpret because B. fragilis is also widespread among healthy people; the Danish group’s claim is that not all B. fragilis are created equal, and that a previously undocumented phage may mark (or drive) a more pathogenic state.
The study, published in Communications Medicine, analyzed stool samples from 877 individuals across Europe, the United States, and Asia. Colorectal cancer cases were about twice as likely as controls to carry traces of the phage. The authors emphasize—correctly—that this is association, not mechanism. They are now pursuing lab and animal studies to test whether the phage changes bacterial behavior in ways that could plausibly contribute to carcinogenesis.
What makes the finding interesting is not the “virus causes cancer” headline bait, but the hypothesis it implies: that microbial mobile genetic elements (phages, plasmids, transposons) might be the missing causal link between a familiar organism and a heterogeneous clinical outcome. A phage could, in principle, alter bacterial toxin production, immune interactions, mucosal barrier effects, or biofilm formation—any of which could change the inflammatory and genotoxic environment in the colon.
But the same logic also raises the obvious epidemiological hazards. Metagenomic signals are sensitive to batch effects, sequencing depth, geography, diet, medication (especially antibiotics), bowel prep, and the simple fact that cancer itself changes the gut environment. Reverse causation is not a philosophical problem here; it’s a biological default.
If the association holds up under careful adjustment and prospective sampling, it could eventually inform screening: not as a substitute for colonoscopy or stool blood tests, but as an additional risk stratifier—one more variable in the actuarial soup. The temptation, as always, will be to turn a probabilistic biomarker into a deterministic story.
The Independent’s reporting on subacute sclerosing pan-encephalitis (SSPE)—a rare, fatal late complication of measles that can emerge years after infection—offers a sobering contrast. There, the causal chain is established, the intervention is straightforward (MMR vaccination), and the tragedy is that compliance is optional. In microbiome-oncology, by contrast, we are still arguing about what the signal is, before we even get to what to do about it.
One story is about a known pathogen and a known tool we keep refusing to use; the other is about a newly found passenger in a bacterial genome that might matter, provided it survives the part where science tries to kill it with replication.
